Products

QIAseq 96-Index L (384)
Cat. No. / ID: 333777

QIAseq 12-Index L (48)
Cat. No. / ID: 333764

QIAseq 12-Index I (48)
Cat. No. / ID: 333714

QIAseq 96-Index I Set A (384)
Cat. No. / ID: 333727

QIAseq 96-Index I Set B (384)
Cat. No. / ID: 333737

QIAseq 96-Index I Set C (384)
Cat. No. / ID: 333747

QIAseq 96-Index I Set D (384)
Cat. No. / ID: 333757

QIAseq 8-Unique Dual Index Set A (48)
Cat. No. / ID: 333715

QIAseq 8-Unique Dual Index Set B (48)
Cat. No. / ID: 333716

QIAseq 96-Unique Dual Index Set A (384)
Cat. No. / ID: 333725

QIAseq 96-Unique Dual Index Set B (384)
Cat. No. / ID: 333735
Features
- Optimized design exclusively from QIAGEN
- QC of manufacturing lots by NGS
- Tube and plate formats available for convenience and scalable sample throughput
- Unique Dual Index adapters available for multiplexing up to 192 samples with QIAseq Targeted NGS solutions*
* Not suitable for QIAseq library prep kits that utilize Y-shaped adapter styles. See our QIAseq FX DNA Library Kits for information on QIAseq Unique Dual Index Y-Adapter Kits.
Product Details
QIAseq Targeted Panels enable Sample to Insight next-generation sequencing (NGS) of DNA or RNA. These optimized solutions facilitate ultrasensitive variant and clonotype detection from cells, tissue and biofluids using integrated Unique Molecular Indices (UMIs).
In order to de-multiplex pooled libraries and assign reads to the appropriate original source library, NGS workflows require the addition of an adapter complex that includes both a common nucleotide sequence, as well as sample index oligonucleotides.
Compatible only with Illumina instruments, our Unique Dual Indexes (UDIs) are optimized for QIAseq Targeted Panel workflows, including QIAseq Targeted DNA, QIAseq RNAscan and QIAseq Immune Repertoire RNA Library Kits. Exclusively available from QIAGEN for QIAseq targeted NGS applications, QIAseq UDI Kits provide the highest level of process safety and ensure confidence in sequencing data by mitigating “index hopping” and mis-assignment of reads. The combination of both UMIs and UDIs provide QIAseq Targeted Panels with a remarkable level of accuracy. Combinatorial, non-unique dual index adapters are also available.
Need a quote for your research project or would you like to discuss your project with our specialist team? Contact Us
Performance
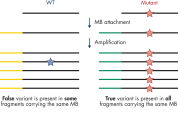
- Accuracy: Innovative digital sequencing (incorporating molecular barcodes) eliminates PCR duplication and amplification artifacts to detect low-frequency variants with high confidence (see figure Principle of molecular barcodes).
- Specificity: The unique combination of our proprietary primer design algorithm and rigorous testing of every primer assay guarantees high specificity and accurate results.
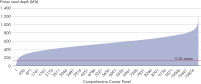
- Uniformity: The QIAseq Targeted DNA Panel workflow has been optimized to deliver highly uniform sequencing results, to ensure sequencing capacity is utilized very efficiently (see figure Uniformity).
- Sensitivity: Digital DNA sequencing approach is optimized to deliver high confidence in calling low-frequency DNA variants. Over 90% sensitivity for 1% NA12878 SNP and indel on typical coding region with false positive less than 15 per mega base region when variants are detected with tiled primer design to cover complete coding region of each gene.
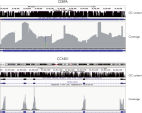
- Universality: The chemistry used in the QIAseq Targeted DNA Panels and workflow is compatible with both regular and GC-rich genomic regions, allowing one to achieve 100% coverage of genes rich in GC content such as CEBPA and CCND1 (see figure Coverage of GC-rich genomic regions).
- Flexibility: The QIAseq Targeted DNA Panels offer a high degree of flexibility in content and sample multiplexing. Several cataloged panels have been developed for a wide range of applications. One can also build a custom panel for a specific content, or extend the contents of an existing cataloged panel. Up to 384 samples can be multiplexed using the QIAseq indices.
See figures
Principle
PCR duplicates are a major issue in targeted DNA sequencing, since, through PCR amplification, they turn unique DNA molecules into identical DNA molecules that cannot be distinguished from each other. In addition, errors from PCR amplification and sequencing process may also be present in final reads that lead to false-positive variants in sequencing results. This, in turn, results in the inability to confidently call DNA variants present at low frequencies in the starting DNA material. To overcome the issue of PCR duplicates and amplification artifacts, the QIAseq Targeted DNA Panels use digital sequencing by incorporating molecular barcodes into the starting DNA material before any amplification takes place, thereby preserving the uniqueness of the starting DNA molecules and overcoming the issues of PCR duplicates, false positives and library bias.
Procedure
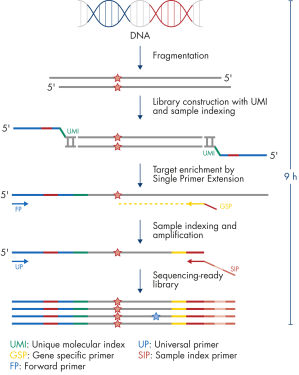
The entire workflow of the QIAseq Targeted DNA Panels to go from extracted DNA to sequencing-ready libraries can be completed in 9 hours (see figure Workflow). Extracted DNA is fragmented, genomic targets are molecularly barcoded and enriched, and libraries are constructed. Sequencing files can be fed into the QIAseq pipeline, a cloud-based data analysis pipeline, which will filter, map and align reads, as well as count unique molecular barcodes associated with targeted genomic regions, and call variants with a barcode-aware algorithm. This data can then be fed into IVA or QCI for interpretation.
See figures
Applications
QIAseq Targeted DNA Indices can be used for indexing samples enriched with either QIAseq Targeted DNA Panels or QIAseq Targeted RNAscan Panels, and are available for both Illumina and Ion Torrent platforms.
Supporting data and figures
Workflow
Isolated DNA, as low as 20 ng, is enzymatically fragmented to generate small pieces of dsDNA. This is followed by the library construction step, in which IL-N7 adapters, molecular barcodes, and sample indexes are incorporated into DNA fragments generated in the previous step. Library fragments now serve as templates for target enrichment using single primer extension. In this step, targets are enriched using a single gene-specific primer and a universal forward primer. The final step is library amplification and sample indexing (for dual indexing) using the IL-S5 sample index primer and a universal primer.