遺伝子発現プロファイリングのための
QIAseq Targeted RNA Panel (12)
Cat. No. / ID: 333002
特徴
- 分子バーコードでPCR duplicate および増幅バイアスを除去
- わずか25 ng のトータルRNA からスタート
- シンプルかつ1日で完了するライブラリー調製ワークフロー
- イルミナおよびサーモフィッシャーシークエンサーに対応
製品詳細
QIAseq Targeted RNA Panelsは、RNAseq による定量的遺伝子発現プロファイリングが可能なプライマーセットおよびライブラリー調製試薬です。分子バーコード技術を2段階PCR ベースの増幅ステップに組み込むことにより、バイアスを最小限に抑えた正確で定量的なRNA シークエンシングを実現します。また、ウェット検証済みのプライマーを用いており、安心してご使用いただけます。
パフォーマンス
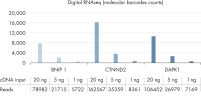
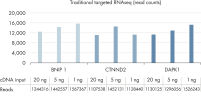
- Accuracy: Innovative digital sequencing (molecular barcode counting) eliminates PCR duplication and amplification bias to deliver the most accurate results (see figure Unbiased and accurate gene quantification) .
- Specificity: The unique combination of our proprietary primer design algorithm and rigorous testing of every primer assay guarantees high specificity and accurate results (see figure Proprietary primer design delivers gene-specific amplicons – 97% specificity).
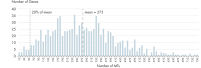
- Uniformity: The QIAseq Targeted RNA Panel workflow has been optimized to deliver highly uniform sequencing results, to ensure sequencing capacity is utilized very efficiently. In fact, digital sequencing (molecular barcodes) entirely remove this variation in RNAseq counting (see figure Unmatched uniformity – 97% of assays are within 20% of median molecular tag counts).
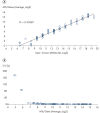
- Reproducibility: The QIASeq Targeted RNA Panel system demonstrates strong correlations across technical replicates, product lots and instruments with correlation coefficients averaging above >0.99, to ensure reliable detection of differences in expression between biological samples
- Sensitivity: Digital RNA sequencing system is optimized to deliver highly reliable quantification down to ~100 copies of an RNA target in 25 ng total RNA (see figure Positive results with as little as 0.2 copies of RNA per cell).
- Flexibility: QIAseq panels combine the power of NGS with the accuracy of qPCR to allow multiplexing of several samples per NGS while delivering cost-effective results (see figure Simple procedure).
図参照
原理
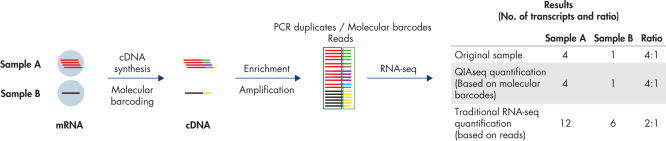
- Traditional RNA sequencing methods suffer from PCR duplication and amplification bias, resulting in inaccurate gene expression analysis. By introducing molecular barcodes before any amplification takes place, QIAseq Targeted RNA Panels are able to eliminate this issue to deliver accurate and digital quantification of genes (see figure Unbiased and accurate gene quantification).
- A unique feature of the QIAseq Targeted RNA Panels is the set of built-in control assays. The gDNA assays control for any gDNA contamination in the RNA sample to ensure reproducible results. The housekeeping gene (HKG) assays are used to normalize data, thereby making sample-to-sample and run-to-run comparisons possible.
図参照
操作手順
- The QIAseq Targeted RNA Panels workflow begins with converting total RNA into cDNA (see figure Simple procedure). The workflow requires minimal RNA input: as little as 25 ng total RNA can be used. No enrichment or depletion steps are necessary. The molecular barcoding step makes use of molecularly barcoded gene-specific primer (GSP1) in a multiplex primer panel (targeting 12-1000 genes) and an input of 20ng of cDNA equivalent (cDNA made from 20 ng of total RNA). After the barcoding step, the uniquely tagged cDNA is purified over beads to remove residual primers, and a PCR is set up with a second pool of gene-specific adapter primers (GSP2) and the RS2 primer, which primes off of a common tag on the GSP1 primers. This reaction insures that intended targets are enriched sufficiently to be represented in the final library. The number of cycles is kept to a minimum to keep PCR-induced variations in amplification to a low level (any variations are easily corrected and accounted for with the molecular barcodes). Another quick cleanup with beads is performed, and a universal PCR is run with RS2 and FS2 primers, which also adds sample-indexing barcodes to each sample. A final cleanup with beads is performed and the library is complete, and ready for quantification and sequencing.
- An integral component of the QIAseq Targeted RNA Panels is data analysis and insight. Data analysis modules have been developed that are comprehensive, yet easy to use. Using these modules require no bioinformatics expertise. Starting with raw reads directly off the sequencer, the QIAseq targeted RNA data analysis tools atQIAGEN’s GeneGlobe portal, provide you with gene counts and fold changes, as well as links for pathway analysis.
図参照
アプリケーション
- Gene expression profiling
- Biomarker research
- Confirmation of whole transcriptome sequencing data
- Confirmation of microarray data
裏付けデータと数値
Digital sequencing (molecular barcodes) principle
QIAseq Targeted RNA Panels use a digital sequencing method, whereby a unique 12-base random molecular barcode incorporated into the gene-specific primers (GSP1) is used in the first extension step (after mRNA is converted to cDNA). Thus, every extension event yields a unique combination of molecular barcode and target sequence. At the end of sequencing, the relative amount of each mRNA target is determined by the number of unique molecular barcode-target combinations that were sequenced, thereby eliminating PCR duplicates and amplification bias, resulting in more accurate, unbiased gene expression analysis.