QuantiTect Probe PCR Kit (200)
Cat. No. / ID: 204343
Features
- Highly sensitive detection of low-copy targets
- qPCR using sequence-specific probes or SYBR Green
- Accurate quantification over several logs of template
- No need to optimize reaction and cycling conditions
- Detection of up to 5 targets in the same tube
Product Details
QuantiTect PCR Kits (qPCR kits) enable sensitive quantification of gDNA and cDNA targets by real-time PCR and two-step RT-PCR using sequence-specific probes or SYBR Green I detection. The real-time PCR kits also allow reliable quantification of up to 5 gDNA or cDNA targets in a single tube by multiplex, real-time PCR or two-step RT-PCR. The combination of a hot start and a unique PCR buffer system in the ready-to-use master mix ensures highly sensitive qPCR on any real-time cycler without the need for optimization. The dNTP mix includes dUTP, allowing optional treatment with UNG. For convenience, the master mixes in QuantiTect PCR Kits can be stored at 2–8°C.
Two kit formats are available for multiplex PCR using sequence-specific probes: the QuantiTect Multiplex PCR Kit for cyclers that require ROX dye for fluorescence normalization, and the QuantiTect Multiplex PCR NoROX Kit for all other cyclers.
Performance
HotStarTaq DNA Polymerase included in QuantiTect quantitative PCR kits increases the specificity of the PCR reaction by providing the most stringent hot start compared with other polymerases.
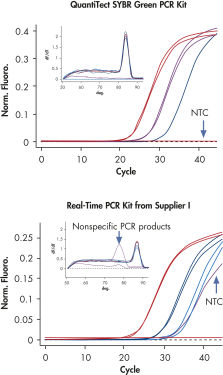
QuantiTect SYBR Green PCR Kits allow specific quantification over a wide linear range (see figure " Specific quantification over a wide linear range"). When used in combination with the QuantiTect Reverse Transcription Kit and QuantiTect Primer Assays, QuantiTect SYBR Green PCR Kits deliver sensitive and reliable results (see figure " Specific and sensitive quantification").
QuantiTect Probe PCR Kits deliver sensitive and reliable results in combination with the QuantiTect Reverse Transcription Kit (see figure " High sensitivity and efficiency, and wide dynamic range"). The unique PCR buffer composition enables QuantiTect Probe PCR Kits to provide sensitive quantification of low-copy DNA targets, as well as accurate quantification over a wide linear range (see figure " Wide dynamic range in real-time PCR").
The QuantiTect Multiplex PCR Master Mix ensures that the PCR products in a multiplex reaction are amplified with the same efficiency and sensitivity as the PCR products in the corresponding single-amplification reactions (see table “Comparable threshold cycle (CΤ) values with triplex and singleplex PCRs”, and figure “ Comparable results with 4-plex and singleplex PCRs”.
| Detection of t(8;14) translocation sequence (20 copies) | Detection of GAPDH cDNA sequence (106 copies) | Detection of NFKB cDNA sequence (see first column for copy number) | |
|---|---|---|---|
| Triplex PCR with 105 copies of NFKB | 34.31 | 20.37 | 21.92 |
| Corresponding singleplex PCRs | 34.07 | 20.54 | 21.83 |
| Triplex PCR with 104 copies of NFKB | 34.61 | 20.62 | 25.03 |
| Corresponding singleplex PCRs | 34.00 | 20.46 | 25.19 |
| Triplex PCR with 103 copies of NFKB | 35.17 | 19.94 | 28.38 |
| Corresponding singleplex PCRs | 34.43 | 20.50 | 28.65 |
As few as 10 copies of a target gene can be detected with the kits, even if the copy number of the reference gene in the same reaction is up to 106-fold greater (see figure “ Detection of 10 copies of target gene with excess reference gene" and the corresponding table "Successful quantitative triplex qPCR of low and high abundance targets").
| Detection of CSBG | Detection of GAPDH | Detection of HSP | |
| Template mix 1 | 1400 copies | 106 copies | 2 x 104 copies |
|---|---|---|---|
| CT values for triplex PCR | 27.73 | 18.69 | 23.59 |
| CT values for singleplex PCRs | 27.08 | 18.89 | 23.52 |
| Template mix 2 | 140 copies | 106 copies | 2 x 103 copies |
| CT values for triplex PCR | 31.11 | 19.00 | 27.05 |
| CT values for singleplex PCRs | 30.66 | 18.61 | 26.97 |
| Template mix 3 | 14 copies | 106 copies | 2 x 102 copies |
| CT values for triplex PCR | 34.74 | 18.98 | 30.71 |
| CT values for singleplex PCRs | 33.84 | 19.01 | 30.38 |
See figures
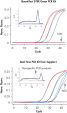 Specific and sensitive quantification.
Specific and sensitive quantification. High sensitivity and efficiency, and wide dynamic range.
High sensitivity and efficiency, and wide dynamic range.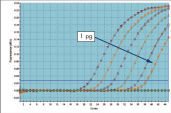 Wide dynamic range in real-time PCR.
Wide dynamic range in real-time PCR.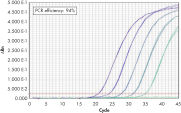 Comparable results between 4-plex PCR and corresponding singleplex PCRs.
Comparable results between 4-plex PCR and corresponding singleplex PCRs.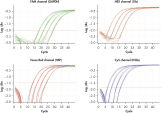 Detection of 10 copies of target gene with excess reference gene.
Detection of 10 copies of target gene with excess reference gene.
Principle
QuantiTect SYBR Green PCR Kits contain an optimized, ready-to-use master mix for highly specific and sensitive real-time quantification cDNA targets using SYBR Green I (see table “Components of 2x QuantiTect SYBR Green PCR Kit”). The fluorescent dye SYBR Green I in the master mix enables the analysis of many different targets without having to synthesize target-specific labeled probes. A balanced combination of K+ and NH4+ ions in the PCR buffer promotes specific primer annealing and enables high PCR specificity and sensitivity (see figure "Specific primer annealing"). In addition, HotStarTaq DNA Polymerase provides a stringent hot start, preventing the formation of nonspecific products.
| Component | Features | Benefits |
| HotStarTaq DNA Polymerase | 15 min activation at 95ºC | Set-up of qPCR reactions at room temperature |
| QuantiTect SYBR Green PCR Buffer | Balanced combination of NH4+ and K+ ions | Specific primer annealing ensures reliable PCR results |
| dNTP mix | Includes dUTP, which partially replaces dTTP and enables optional UNG treatment of reactions | Eliminates contamination from carryover of PCR products by optional UNG treatment |
| SYBR Green I dye | Yields a strong fluorescent signal upon binding double-stranded DNA | Highly sensitive quantification |
| ROX dye | For normalization of fluorescent signals on Applied Biosystems and, optionally, Agilent instruments | Precise quantification on cyclers that require ROX dye. Does not interfere with reactions on other real-time cyclers |
QuantiTect Probe PCR Kits contain an optimized, ready-to-use master mix for highly specific and sensitive real-time quantification of gDNA and cDNA targets using sequence-specific probes (see table “Components of 2x QuantiTect Probe PCR Kit”). The kits are designed for use with all types of sequence-specific probes, including hydrolysis probe detection (e.g., TaqMan® and other dual-labeled probes), FRET probes, and Molecular Beacons. QuantiTect Probe PCR Kits contain a unique PCR buffer that contains a balanced combination of K+ and NH4+ ions, which promote specific primer annealing, enabling high PCR specificity and sensitivity (see figure "Specific primer annealing"). In addition, HotStarTaq DNA Polymerase provides a stringent hot start, preventing the formation of nonspecific products.
| Component | Features | Benefits |
| HotStarTaq DNA Polymerase | 15 min activation at 95ºC | Set-up of qPCR reactions at room temperature |
| QuantiTect Probe PCR Buffer | Balanced combination of NH4+ and K+ ions | Specific primer annealing ensures reliable PCR results |
| dNTP mix | Includes dUTP, which partially replaces dTTP and enables optional UNG treatment of reactions | Eliminates contamination from carryover of PCR products by optional UNG treatment |
| ROX dye | For normalization of fluorescent signals on Applied Biosystems and, optionally, Agilent instruments | Precise quantification on cyclers that require ROX dye. Does not interfere with reactions on other real-time cyclers |
QuantiTect Multiplex PCR Kits enable success in multiplex, two-step RT-PCR on the first attempt (see flowchart " QIAGEN multiplex kits"). The optimized master mix ensures that PCR products in a multiplex reaction are amplified with the same efficiency and sensitivity as PCR products in a corresponding single-amplification reactions. As few as 10 copies of a target gene can be detected with the kit.
Amplifying reference and target genes in the same reaction instead of in separate reactions increases the reliability of gene quantification by minimizing handling errors. The QuantiTect Multiplex PCR Master Mix contains a balanced combination of K+ and NH4+ ions as well as the unique synthetic Factor MP, which together promote stable and efficient annealing of primers and probes to the nucleic acid template, enabling high PCR efficiency (see table "Components of 2x QuantiTect Multiplex PCR Kit"). In addition, HotStarTaq DNA Polymerase provides a stringent hot start, preventing the formation of nonspecific products.
The master mixes in QuantiTect PCR Kits also contain dUTP, enabling pretreatment with uracil-N-glycosylase (UNG) prior to starting PCR, which ensures that any contaminating PCR products do not affect subsequent PCR reactions.
| Component | Features | Benefits |
| HotStarTaq DNA Polymerase | 15 min activation at 95ºC | Set-up of qPCR reactions at room temperature |
| QuantiTect Multiplex PCR Buffer | Balanced combination of NH4+ and K+ ions | Specific primer annealing ensures reliable PCR results |
| Synthetic Factor MP | Reliable multiplexing analysis of up to 4 genes in the same tube | |
| dNTP mix | Includes dUTP, which partially replaces dTTP and enables optional UNG treatment of reactions | Eliminates contamination from carryover of PCR products by optional UNG treatment |
| ROX dye* | Normalizes fluorescent signals on Applied Biosystems and, optionally, Agilent instruments | Precise quantification on cyclers that require ROX dye. Does not interfere with reactions on other real-time cyclers |
See figures
Procedure
If required, reactions can be pretreated with uracil-N-glycosylase (UNG) to eliminate carryover of PCR products from previous reactions. For optimal results in real-time two-step RT-PCR, we recommend synthesizing cDNA using the QuantiTect Reverse Transcription Kit. The kit provides fast cDNA synthesis in just 20 minutes with integrated removal of genomic DNA contamination.
QuantiTect SYBR Green and Probe PCR Kits overcome the need for optimization of reaction conditions, which can be tedious and time-consuming. Simply add primers and DNA template to the ready-to-use PCR master mix, and start the reaction. Follow the protocol in the handbook to get fast and reliable results on any real-time cycler.
Highly specific results in gene expression analysis are guaranteed when QuantiTect SYBR Green PCR Kits are used in combination with QuantiTect Primer Assays. These are genomewide, bioinformatically validated primer sets for detecting transcripts from human, mouse, rat, and many other species. QuantiTect Primer Assays can be easily ordered online at GeneGlobe.
QuantiTect Multiplex PCR Kits contain ready-to-use master mixes that eliminate the need for optimization of reaction and cycling conditions. The handbook contains a single protocol that can be used with all available real-time cyclers and also lists recommended dyes.
Kits are available with or without ROX passive reference dye in the master mix, enabling use on virtually any real-time cycler (see table “Choosing the right QuantiTect Multiplex PCR Kit”). Due to the optimized ROX concentrations, detection of even low copy numbers is achieved through automatic data analysis.
| ROX dye | Kit | Compatible cyclers |
| Supplied in master mix | QuantiTect Multiplex PCR Kit | Cyclers from Applied Biosystems |
| Absent from master mix | QuantiTect Multiplex PCR NoROX Kit | Rotor-Gene cyclers, and cyclers from Bio-Rad, Cepheid, Eppendorf, Roche, Agilent, and other suppliers |
Applications
QuantiTect PCR Kits can be used for gene expression analysis of cDNA or quantification of gDNA on any real-time cycler. This includes instruments from Applied Biosystems, Bio-Rad, Cepheid, Eppendorf, Roche and Agilent. For the Rotor-Gene Q and other Rotor-Gene cyclers, we recommend using the Rotor-Gene SYBR Green PCR Kit, Rotor-Gene Probe PCR Kit or Rotor-Gene Multiplex PCR Kit, which have been specially developed for fast cycling on these instruments.
| Features | QuantiTect SYBR Green PCR Kits | QuantiTect Probe PCR Kits | QuantiTect Multiplex PCR Kits |
| Applications | Real-time quantification of genomic DNA or cDNA targets | Real-time quantification of genomic DNA or cDNA targets | Real-time quantification of genomic DNA or cDNA targets |
| Reaction type | PCR and two-step RT-PCR | PCR and two-step RT-PCR | Multiplex PCR and multiplex two-step RT-PCR |
| Real-time or endpoint | Real-time | Real-time | Real-time |
| Sample/target type | DNA, cDNA | DNA, cDNA | DNA, cDNA |
| Single or multiplex | Single | Single | Multiplex |
| SYBR Green I or sequence-specific probes | SYBR Green I | Sequence-specific probes | Sequence-specific probes |
| Thermal cycler | All real-time cyclers (e.g., LightCycler, Rotor-Gene, ABI) | Most real-time cyclers (e.g., LightCycler, Rotor-Gene, ABI) | Most real-time cyclers (e.g., LightCycler, Rotor-Gene, ABI) |
| With or without ROX | With ROX | With ROX | With or without ROX |
Supporting data and figures
Specific quantification over a wide linear range.