DNeasy Plant Pro Kit (50)
Cat. No. / ID: 69204
Features
- Superior PCR performance with patented Inhibitor Removal Technology (IRT)
- Rapid extraction of ready-to-use DNA
- No organic extraction, no ethanol precipitation
- Highly efficient lysis and release of DNA from tough plant materials and associated plant pathogens
Product Details
The DNeasy Plant Pro Kit has several innovative features to overcome the challenges of DNA extraction from plant tissue and enable recovery of high-quality DNA from the toughest sample types, including strawberry leaf, grapevine leaf, pine needles and various seed types. Increased yields of plant DNA and plant pathogen DNA combined with superior removal of inhibitors ensure high-performance results in sensitive downstream applications.
Performance
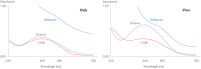
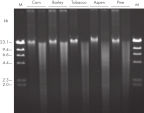
The DNeasy Plant Pro Kit – the newest member of the trusted DNeasy Plant family – enables purification of significantly higher yields of DNA from the toughest sample types, including strawberry leaf, grapevine leaf, pine needles and various seed types (see table "Selection of plant species processed with DNeasy Kits" and see figure “ Significantly higher yields of pure DNA”). Patented Inhibitor Removal Technology (IRT) provides superior removal of inhibitors without using harsh chemicals, yielding pure DNA with less PCR inhibition (see figure “ Efficient removal of PCR inhibitors”).
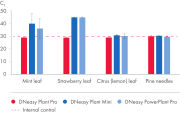
When CT values of PCR reactions with plant DNA eluates containing possible inhibitors were compared to CT values of the PCR reaction with water added as control which does not inhibit amplification of the IC DNA, the eluate from the DNeasy Plant Pro Kit showed no inhibition.
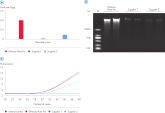
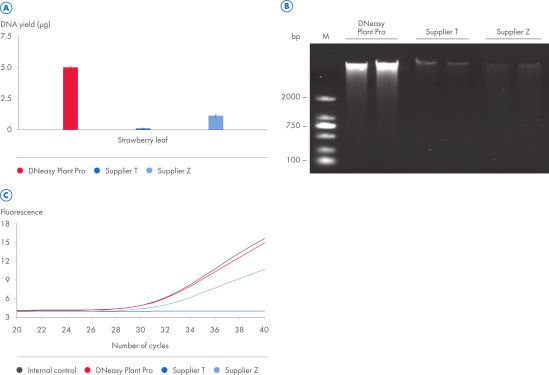
With the DNeasy Plant Pro Kit, yields of DNA purified from strawberry leaf – a particularly tough plant sample – were significantly higher than those obtained using a kit from Supplier T and Supplier Z. Furthermore, the DNeasy Plant Pro Kit provided superior removal of inhibitors and the DNA eluate showed no inhibition (see figure “ Significantly higher yields and superior inhibitor removal”). The DNeasy Plant Pro Kit can also purify bacterial, fungal and viral DNA from plant and root samples.
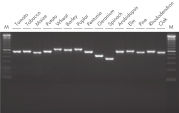
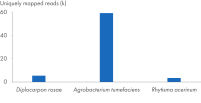
The kit can be successfully combined into a workflow with proven products optimized for next-generation sequencing (NGS) (see figure “ Optimized NGS workflow”). Rapid and reliable identification of plant pathogens is crucial to avoid disease spread and facilitate effective disease management. Plant and pathogen DNA co-purified using the DNeasy Plant Pro Kit enables successful identification of a range of pathogens (see figure “ Successful identification of plant pathogens by NGS”)
| Abies alba (silver fir) | Nicotiana tabacum (tobacco) |
| Aesculus hippocastanum (horse chestnut) | Oryza sativa (rice)4 |
| Arabidopsis thaliana (thale cress) | Pelargonium sp. (geranium)4 |
| Avena sp. (oat) | Petunia sp.4 |
| Brassica napus (oilseed rape) | Pinus sylvestris (Scotch pine), P. brutia5 |
| Brassica oleracea (kohlrabi) | Populus tremula (aspen) |
| Chicorium endivia (chicory) | Pseudotsuga menziesii (Douglas fir) |
| Citrullini lanatus (water melon) | Quercus robur, Q. petrea (oak)6,7 |
| Egeria sp. | Rhododendron sp.2,4 |
| Fagus sylvatica (beech)1 | Rubus idaeus (raspberry) |
| Helianthus spp. (sunflower) | Solanum tuberosum (potato) |
| Hordeum vulgare (barley)2 | Sphagnum palustre (moss) |
| Humulus sp. (hops) | Spinacia oleracea (spinach) |
| Hydrilla sp. | Taxus baccata (yew) |
| Kalanchoe spp. | Triticum aestivum (wheat)4 |
| Lupinus sp. | Ulmus glabra (elm)6 |
| Lycopersicon esculentum (tomato)3 | Vitis spp. (grape)6 |
| Myriophyllum sp. | Zea mays (maize) |
See figures
Principle
The DNeasy Plant Pro Kit comprises a novel and proprietary method for fast and easy purification of total cellular DNA from plant cells, tissues and seeds. In addition, this kit can be used to purify bacterial, fungal and viral DNA from plant and root samples. The DNeasy Plant Pro Kit uses bead-beating technology, which replaces cumbersome DNA isolation procedures such as CTAB, phenol or chloroform extraction to recover high-quality DNA from the toughest sample types, including strawberry leaf, grapevine leaf, pine needles and various seed types. The DNeasy Plant Pro Kit uses the second generation of QIAGEN’s patented Inhibitor Removal Technology (IRT) to remove PCR inhibitors, including polysaccharides, polyphenolics and other secondary metabolites from plant extracts during the isolation process. Improved IRT, combined with efficient bead beating and lysis chemistry, results in high yields of inhibitor-free DNA that is ready for immediate use in downstream applications, including PCR, qPCR, and RAPD analysis, RFLP analysis, Southern blotting and next-generation sequencing.
Procedure
The DNeasy Plan Pro Kit uses bead-beating technology, whch replaces cumbersome DNA isolation procedures to recover high-quality DNA from the toughest sample types. Samples are added to the Tissue Disruption Tube which contains a specially shaped bead and a buffer for rapid homogenization (see figure " Rapid homogenization in Tissue Disruption Tubes"). Cell lysis and DNA release occur by mechanical and chemical methods. Released genomic DNA is cleared of PCR inhibitors using QIAGEN’s second-generation Inhibitor Removal Technology (IRT) and then captured on a silica membrane in a spin column format. DNA is then washed and eluted from the membrane and is ready for PCR, qPCR, NGS and other downstream applications.
An optional Phenolic Separation Solution step helps provide pure DNA even from samples with high polyphenolic compounds, such as pine or grape leaves by breaking the bonds between DNA and phenolics. This step prevents DNA loss that would otherwise occur in these samples.
The DNeasy Plant Pro Kit is used with a vortexer or high-velocity bead beating, based on sample needs. Vortex methods work with soft leaf tissue. High-velocity bead beaters, like the PowerLyzer 24 Homogenizer or the TissueLyser II, break down tougher plant material such as roots, seeds, stems or challenging leaf tissues. Furthermore, purification of DNA using the DNeasy Plant Pro Kit can be fully automated on the QIAcube Connect.
See figures
Applications
DNeasy Plant Kits provide purification of ready-to-use DNA from plant samples, including plant cells, plant tissues and fungi.
The DNeasy Plant Pro Kit is designed for:
- DNA extraction from plants and plant-associated microorganisms
- PCR and NGS analysis
- Marker-assisted breeding
- Plant pathogen research
- Studies on genetically modified plants
- Detection of resistance traits
| Features | DNeasy Plant Mini Kit | DNeasy Plant Maxi Kit | DNAeasy 96 Plant Kit | DNeasy Plant Pro Kit |
|---|---|---|---|---|
| Applications | PCR, qPCR, blotting, next-generation sequencing | PCR, qPCR, blotting, next-generation sequencing | PCR, qPCR, blotting, next-generation sequencing | PCR, qPCR, blotting, next-generation sequencing |
| Elution volume | 50–200 µl | 500 µl – 2 ml | 100–200 µl | 50–100 µl |
| Format | Spin column | Spin column | 96-well plate | Spin column |
| Main sample type | Plant samples | Plant samples | Plant samples | Plant samples and seeds |
| Bead size | N.A. | N.A. | N.A. | 5/32” (3.9 mm) ballcone |
| Binding capacity | Up to 50 µg | Up to 500 µg | Up to 50 µg (per well) | Up to 50 µg |
| Processing | Manual | Manual | Manual | Bead beating |
| Purification of total RNA, miRNA, poly A+ mRNA, DNA or protein | DNA | DNA | DNA | DNA |
| Sample amount | Up to 100 mg | Up to 1 g | Up to 50 mg | Up to 100 mg |
| Technology | Silica technology | Silica technology | Silica technology | Silica technology |
| Throughput | Varies | Varies | 96 or 192 samples | Varies |
| Time per run or per prep | <1 hour | <2 hours | <2 hours (192 samples) | 45 minutes (24 samples) |
| Typical yield (from 50 mg starting material) | Up to 30 µg | Up to 260 µg | Up to 15 µg | Up to 30 µg |
N.A. = Not applicable
Supporting data and figures
Significantly higher yields and superior inhibitor removal.