Taq DNA Polymerase (250 U)
Cat. No. / ID: 201203
Features
- QIAGEN PCR Buffer for minimal optimization
- Additional ready-to-load PCR buffer for faster handling
- Q-Solution for amplification of GC-rich templates
- Choice of formats for convenience and ease of handling
Product Details
Performance
Taq DNA Polymerase outperformed kits tested from other suppliers and delivers robust PCR performance in a wide range of PCR conditions, without the need for time-consuming optimization (see figures " Tolerance of different primer Tm Values" and " Specific amplification of long PCR products"). Every lot of Taq DNA Polymerase is subjected to a comprehensive range of quality control tests, including a stringent PCR specificity and reproducibility assay in which low-copy targets are amplified from human genomic DNA (see figure " Lot-to-lot reproducibility"). The unique formulation of QIAGEN PCR Buffer and CoralLoad PCR Buffer, also provided with the kit, enable highly specific PCR in a variety of PCR conditions with minimal optimization requirements (see figure " Wide annealing-temperature window and Tolerance to variable magnesium concentration"). In addition, CoralLoad PCR Buffer enables immediate loading of PCR products onto an agarose gel for even easier handling and faster results. Suboptimal PCR can be improved using Q-Solution, a PCR additive, also provided with the kit (see figure " Amplification of difficult templates").
Taq DNA Polymerase specifications
Concentration: 5 units/µl
Recombinant enzyme: Yes
Substrate analogs: dNTP, ddNTP, dUTP, biotin-11-dUTP, DIG-11-dUTP, fluorescent-dNTP/ddNTP
Extension rate: 2–4 kb/min at 72°C
Half-life: 10 min at 97°C; 60 min at 94°C
Amplification efficiency: ≥105 fold
5'–>3' exonuclease activity: Yes
Extra A addition: Yes
3'–>5' exonuclease activity: No
Contaminating nucleases: No
Contaminating RNases: No
Contaminating proteases: No
Self-priming activity: No >
See figures
Principle
Taq DNA Polymerase is a high-quality recombinant enzyme that is suitable for general and specialized PCR applications (see figures " Tolerance of different primer Tm Values" and " Specific amplification of long PCR products").
QIAGEN PCR Buffer
Innovative QIAGEN PCR Buffer has been developed to save time and effort by reducing the need for PCR optimization. QIAGEN PCR Buffer contains both KCl and (NH4)2SO4(see figure " Increased specificity of primer annealing"). This unique buffer facilitates the amplification of specific PCR products. During the annealing step of every PCR cycle, the buffer allows a high ratio of specific-to-nonspecific primer binding. Owing to a uniquely balanced combination of KCl and (NH4)2SO4, the PCR buffer provides stringent primer-annealing conditions over a wider range of annealing temperatures and Mg2+ concentrations than conventional PCR buffers. Optimization of PCR by varying the annealing temperature or the Mg2+ concentration is dramatically reduced and often not required (see figure " Wide annealing temperature window and Tolerance of variable magnesium concentration").
CoralLoad PCR Buffer
CoralLoad PCR Buffer has all the advantages of QIAGEN PCR Buffer. In addition, it can also be used to directly load the PCR reaction onto an agarose gel — separate addition of a gel loading buffer is not required. CoralLoad PCR Buffer provides the same high PCR specificity and minimal reaction optimization as the conventional QIAGEN PCR Buffer. Additionally, it contains two marker dyes — an orange dye and a red dye — that facilitate estimation of DNA migration distance and optimization of agarose gel run time (see figure " CoralLoad PCR Buffer"). The buffer ensures improved pipetting visibility and enables direct loading of PCR products onto a gel, for enhanced convenience.
Q-Solution
Q-Solution facilitates amplification of GC-rich templates or templates with a high degree of secondary structure by modifying the melting behavior of DNA. Use of this unique reagent often enables or improves suboptimal PCR (see figure " Amplification of difficult templates"). Unlike DMSO and other PCR additives, Q-Solution is used at a defined working concentration with any primer–template system and is not toxic.
See figures
 Specific amplification of long PCR products.
Specific amplification of long PCR products.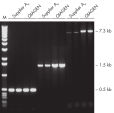 NH4+ and K+ cations in QIAGEN PCR buffers increase specific primer annealing.
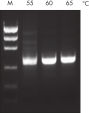
NH4+ and K+ cations in QIAGEN PCR buffers increase specific primer annealing.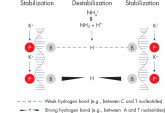 A. Wide annealing temperature window. B. Tolerance of variable magnesium concentration.
A. Wide annealing temperature window. B. Tolerance of variable magnesium concentration.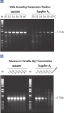 CoralLoad PCR Buffer.
CoralLoad PCR Buffer.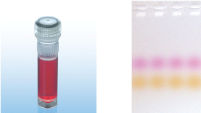 Amplification of difficult templates with Q-Solution.
Amplification of difficult templates with Q-Solution.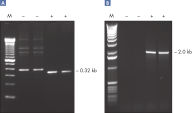
Procedure
Applications
Taq DNA Polymerase is used for standard and specialized applications, including:
- General PCR
- RT-PCR
- Screening
- PCR-based DNA fingerprinting (VNTR, STR, and RAPD)
Supporting data and figures
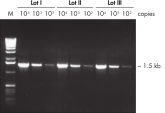
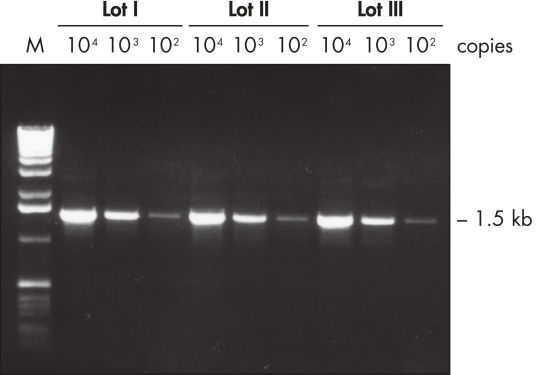
Lot-to-lot reproducibility.
A fragment of the single-copy gene for cystic fibrosis was amplified from 30 ng, 3 ng, and 300 pg human genomic DNA corresponding to 104, 103, and 102 copies of target template, respectively. Three different lots of QIAGEN Taq DNA Polymerase were used and equal volumes of the PCR product were analyzed on a 1% agarose gel. M: markers.
Specifications
| Features | Specifications |
|---|---|
| Applications | PCR, RT-PCR, DNA fingerprinting |
| dNTP's included | No |
| Real-time or endpoint | Endpoint |
| Reaction type | PCR amplification |
| Single or multiplex | Single |
| With/without hotstart | Without hotstart |
| Enzyme activity | 5' -> 3' exonuclease activity |
| Mastermix | No |
| Sample/target type | Genomic DNA and cDNA |